A large international collaboration, led by A/Prof Xiu Ying Wang and Prof Manuel Graeber of the University of Sydney, has developed an innovative, advanced artificial intelligence (AI) application, PathoFusion, that could be used for the examination of routine tissue samples in order to identify indications of cancer.
The research melds contributions from computer scientists, neuropathologists, neuosurgeons, medical oncologists and medical imaging scientists.
ANSTO’s Prof Richard Banati, a Professor of Medical Radiation Sciences/Medical Imaging, who studies the brain’s innate immune system using advanced medical imaging techniques, is a co-author on the paper published in the journal, Cancers.
“The idea behind PathoFusion was to create a novel advanced deep learning model to recognise malignant features and immune response markers, independent of human intervention, and map them simultaneously in a digital image,” explained Banati.
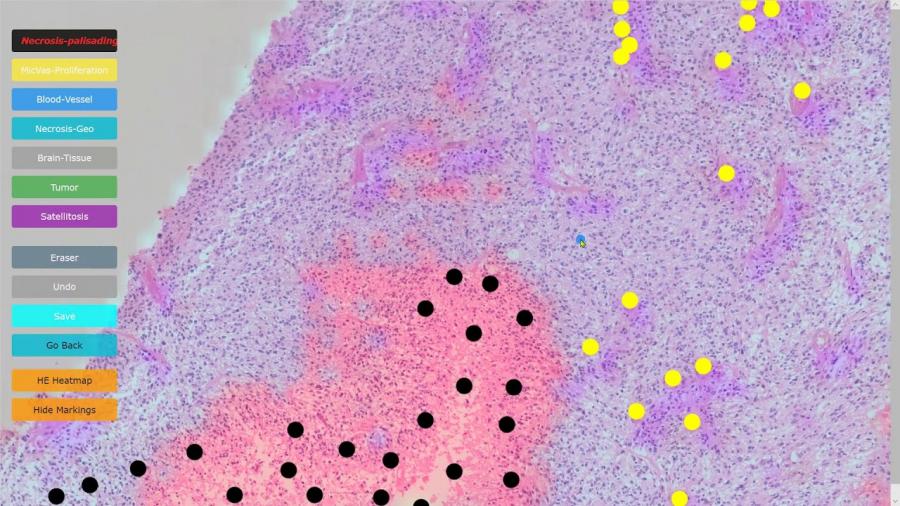
Demonstration of PathoFusion system for automated detection of cancerous features in whole-slide H&E images and mapping with immunohistochemical data in adjacent tissue sections. Courtesy of the University of Sydney
Scientists specifically designed a bifocal deep learning framework which is analogous to how a microscopist works in histopathology image analysis. The framework uses a convolutional neural network (ConvNet/CNN), which was originally developed for natural image classification.
This deep learning algorithm can take in an input image, assign importance to various aspects/objects in the image and differentiate one from another.
The experiment to evaluate the model involved the examination of tissue from cases of glioblastoma, an aggressive cancer that affects the brain or spine.
The team used the expert input of neuropathologists to ‘train’ the software to mark key features.
Experiments confirmed that the application achieved a high level of accuracy in recognising and mapping six typical neuropathological features that are markers of a malignancy.
Pathofusion reliably identified forms and structural features with a precision of 94% and sensitivity of 94.7%. and an immune markers at a precision of 96.2% and sensitivity of 96.1%.

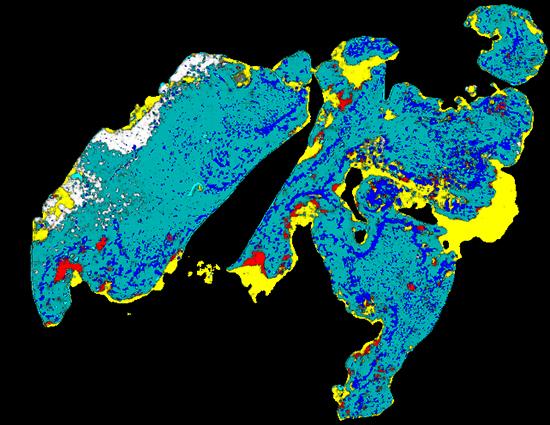
The digital image is a heatmap that illustrates cancerous structures in different colours
The application combines layers of information about dead or dying tissue, the proliferation of microscopic blood vessels and other vasculature with the expression of a tumour genetic marker, CD276, in an image that combines the data in a heatmap.
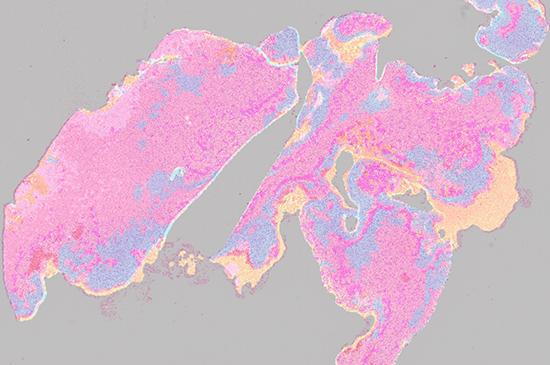
A fusion heatmap of cancerous structures in H&E image and immunopositivity of CD276 marker
The image uses strong colours to depict the features and their distribution. Conventional staining techniques are often monochromatic.
“The research confirmed that it is possible to train neural networks effectively using only a relatively small number of cases, that should be useful for some scenarios,” said Banati.
The research was successful in efficiently training a convolutional neural network to recognise key features in stained slides, improving the model and increasing the effectiveness of feature recognition (with fewer physical cases than conventionally needed for neural network training) and establishing a method to include immunological data.
Painstaking and time-consuming routine morphological diagnostic work is carried out by pathologists, who examine individual slides under a microscope to mark and quantify features that are markers of disease and provide the information to clinicians.
"Anticipating further hardware improvements in computing, it should exceed the speed of human microscopic feature recognition by orders of magnitude when whole histological slides are used at high resolution," said Prof Manuel Graeber, Barnet-Cropper Chair of Brain Tumour Research, The University of Sydney and Brain and Mind Centre.
“In future, the model could improve the workflow of a neuropathology or pathology unit by facilitating microscopic analyses or benefit patients in area without local access to these services, “ explained Banati.
Contributors to the research included The University of Sydney, The Brain and Mind Centre, Xuanwu Hospital (China), St Vincent’s Hospital, Cooperative Trials Group of Neuro-Oncology, Brain Cancer Consultancy, University of Western Australia