In a continuation of pioneering work on food provenance, ANSTO researchers have developed a universal approach to identify the origin of marine animals on a global scale, which could help combat food fraud.
In research published in Fish and Fisheries last week, lead author Dr Jasmin Martino of ANSTO and associates at ANSTO, the University of South Australia and the University of Southampton, demonstrated how universal chemical markers and isoscape mapping of ocean areas can successfully geolocate a diverse range of seafood back to the region of origin.
These techniques have the potential to change the way the provenance of seafood is verified on a more unified, global, and equitable scale.

The method, which is based on the use of stable oxygen isotopes as universal markers, was tested on a range of seafood from diverse geographical locations.
The team used natural chemical markers imprinted within the shells and bones of marine animals that permanently retain lifetime chemical fingerprints and do not decay.
Because these markers reflect the animal’s aquatic environment, they can be used to locate geographic origin.
“At this stage of the research, this is most applicable to broad-scale comparisons, such as tropical versus temperate waters, where we can isolate predictable environmental variation,” explained Dr Martino.
This study employed a universal chemical marker that is expressed similarly among different marine animals within the same geographical region.
These universal chemical markers called oxygen isotope values, specifically the ratio between oxygen atoms of different masses, are precisely controlled by ocean composition and temperature rather than an animal’s biology.
Harnessing the predictable spatial variation of marine habitats, the researchers were able to construct a global ocean map, called an isoscape, that predicts the oxygen isotope imprint in shells and bones across species.
The accuracy of the predictive isoscape was evaluated against a database of authentic, geo-referenced oxygen isotope values from the literature (3954 data points representing 68 species).
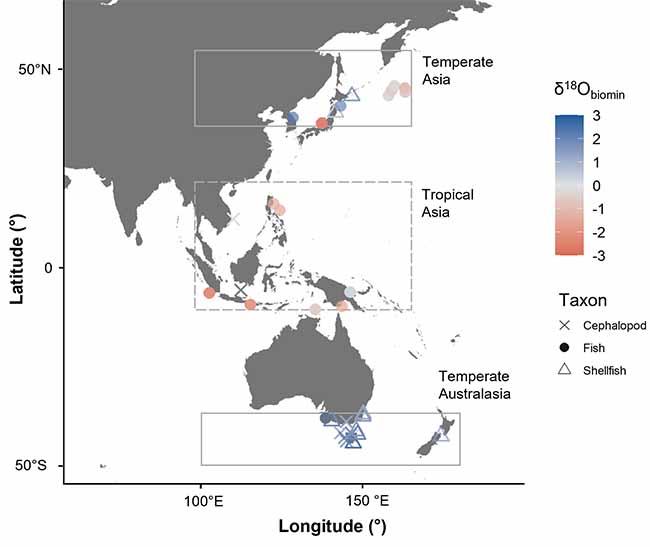
The universal framework was then rigorously tested with a machine learning model approach developed at ANSTO, which confirmed that the global isoscape could be used to correctly identify the origins of a wide range of marine animals living in different latitudes with up to 90 per cent accuracy.
Fish, cephalopods, such as squid and octopus, and shellfish found between the tropical waters of southeast Asia and the cooler waters of southern Australia were classified correctly.
The next step will be the integration of oxygen isotopes with additional universal markers to capture longitude or smaller spatial scales and further refine the approach.
“We have a few promising candidate markers in mind that will help with improving spatial resolution,” said Dr Martino.
Other ANSTO scientists who contributed to this research included Dr Debashish Mazumder, Leader of Food Provenance project and Dr Jagoda Crawford.
ANSTO is collaborating with universities, government organisations and industries in Australia on seafood provenance research to combat food fraud and ensure integrity in the seafood supply chain.
Read more on The Conversation






